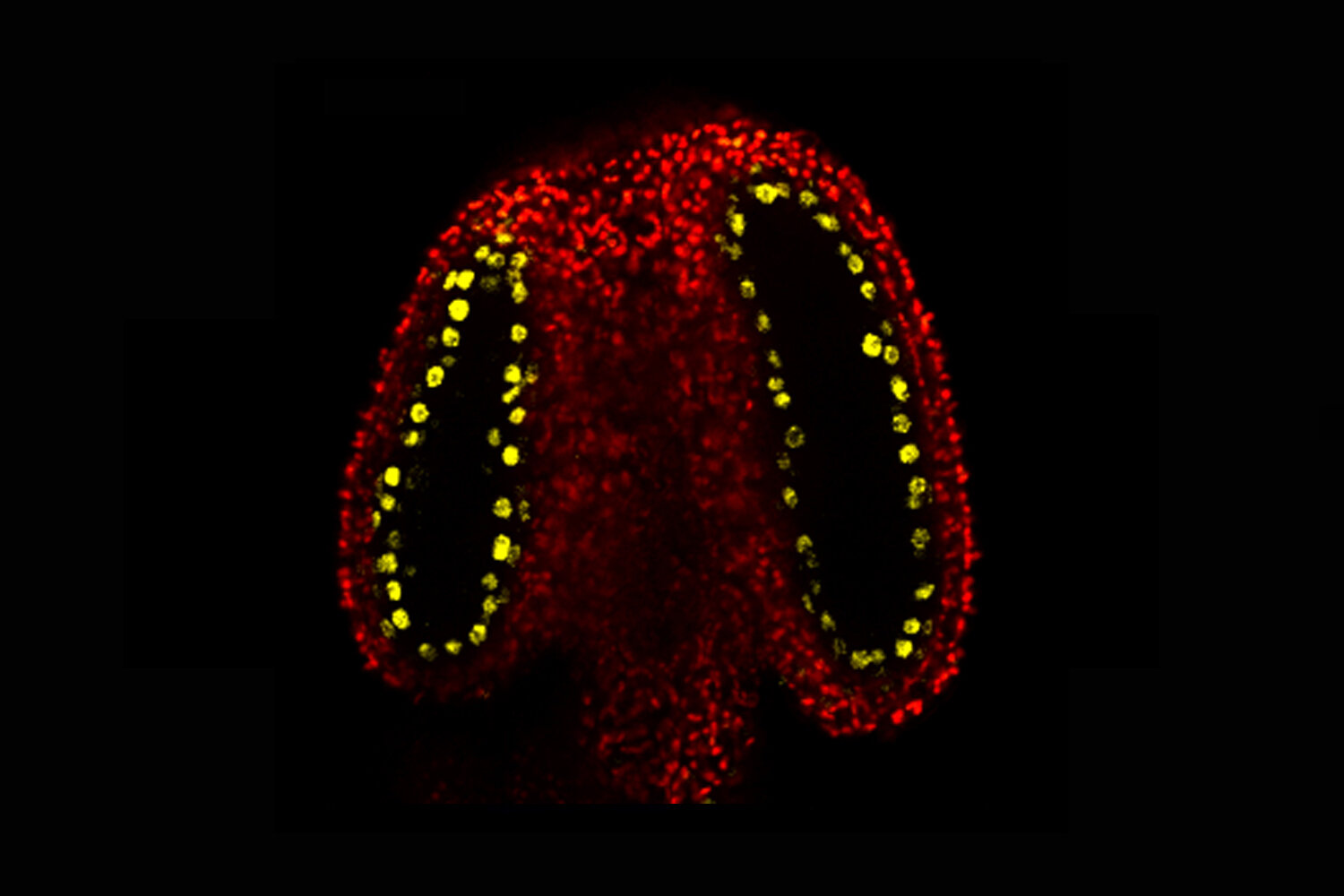
CLSY3 is a yellow fluorescent protein fused with. It is expressed specifically in the tapetal cells around the germ cells. Credit: John Innes CenterThe genetic code, DNA, passes hereditary information from parent to child. Epigenetically, chemically induced changes around the DNA can also pass this information.The John Innes Centre's new research has revealed a mechanism that adjusts these modifications and alters the way information is passed down through the generations.DNA methylation is one example of epigenetic modifications. It occurs when a chemical cap or methyl group is added to DNA. This switches on or off a gene or set of genes.Some methyl markers in germline cells (eggs, sperm) are reset as they develop. This affects the information that is passed on to the next generation.It is not clear how this happens during plant reproduction.Science published an exciting study that reveals the molecular mechanism for DNA methylation and reprogramming in male plants' germline.The anthers, which are the male reproductive parts of the plant, contain cells that divide to make sperm. They are surrounded with cells that nourish them. These nurse cells are known as tapetal cells.The John Innes Centre team found that tapetal cells produced a lot of small RNA molecules. They discovered this was caused by CLSY3, a protein that is located in tapetal cells in the anther. These small RNAs are found to move from the taptal cells into meiocytes. They add new methyl marks (unstable genetic element) to the transposons.When tapetal smallRNAs are absent, Gypsy1 transposon can be seen as active (shown in yellow fluorescence). Credit: John Innes Center"This discovery has changed the way we view epigenetic inheritance between generations of plants. The sperm's DNA methylome can be determined by small RNAs that are produced in germline nurse cells. These small RNAs play a key role in determining the inherited methylome. This is a sign of convergent functional evolutionary between animal and plant reproduction," Dr. Xiaoqi Feng (corresponding author), group leader at John Innes Centre.This reprogramming prevents transposons from jumping around inside germ cells and preserves the integrity the genome over time.These small RNAs target genes in meiocytes with the same DNA sequences of the source transposons. This helps to control gene expression and facilitate meiosis (a type of cell division that produces sperm).These findings are applicable to all plant and animal kingdoms, and offer a crucial new insight for the global community of epigenetics researchers. Although previous research has demonstrated that rice and maize have similar small RNA tapets, it was not clear why. This study provides new insights into the mechanistic basis of DNA methylation in commercial crops and could lead to biotechnology.Dr. Jincheng Long was the joint first author. He stated that "our study could open up a new avenue for crop biotechnology." By manipulating small RNA directed DNAmethylation in cells, one can directly influence the process of seed formation and breeding.The study is important for fundamental biological reasons, Dr. James Walker, who is the joint first author, explains that "our work demonstrates paternal epigenetic inheritance can be determined by tapetal cells which drive reprogramming on a scale unparalleled in plants."The molecular mechanism that our research revealed pushes our understanding de novo DNAmethylation to the next stage, showing how new marks are established at certain sites in specific cells."Continue reading Study provides new insights into immortal plant cellsAdditional information: Science (2021) "Nurse cells derived small RNAs determine paternal epigenetic inheritance of Arabidopsis." Information from Science: Journal "Nurse cell-derived small RNAs determine paternal epigenetic inheritability in Arabidopsis." (2021). science.sciencemag.org/cgi/doi 11.26/science.abh0556
