There is a question about what is in a name. It seems like a lot for the organisms.
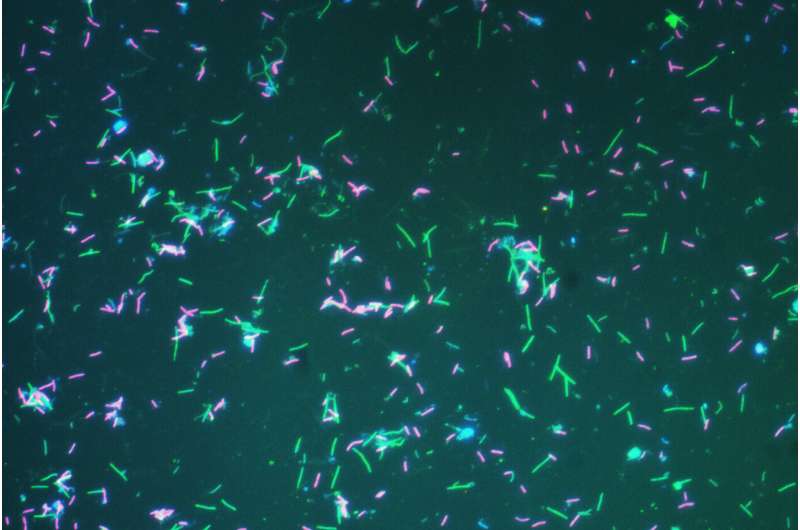
bacteria are an example of prokaryotes that are abundant the world over They can be found in the oceans, soils, hot springs, and other places.
They're everywhere and scientists around the world are trying to communicate about them. The problem is that most don't have a name.
Current regulations of the International Code of Nomenclature of Prokaryotes require new species to be grown in a lab and distributed as pure and viable cultures. To prove it, you have to have more than one specimen.
A team of scientists presented a new system, the SeqCode, and a corresponding registration portal in an article published in the journal Nature Microbiology.
"Our goal is to unify field and laboratory studies in microbiology and respond to significant recent advancements in environmental genomics by providing a path to formally name the majority of identified yet unnamed prokaryotes," said UNLV microbiologist Brian Hedlund. High genome quality standards, good naming practice, and a well-ordered database should be promoted by the SeqCode.
The Seq Code is being created.
More than 850 scientists from more than 40 countries participated in a series of online workshops funded by the National Science Foundation to develop the new SeqCode, which uses genome sequence data for both cultivated and uncultivated prokaryotes as the basis for naming them.
Scientists who study prokaryotes in environments all over the world have used environmentalgenomics techniques to sample and study them, and hundreds of thousands of genomes are available in public databases. An alternative to the ICNP that would accept DNA sequence data and improve resources for researchers was supported by the community that participated in the workshops.
William B. Whitman is a microbiologist at the University of Georgia. The expansion will serve the research and the broader community by providing a common language for all prokaryotes that is organized and supported by data.
To be included in the SeqCode, genomes need to meet rigorous scientific standards. The SeqCode is not universally accepted, but it is in line with international principles for naming other organisms.
Any organisms with a high-quality genome sequence can be named under the SeqCode. All names formed under the ICNP will be accepted by us. The SeqCode will be used more often than the ICNP in the future.
The creation of clarity and chaos.
Authors argue that one of the main goals of the new system is to reverse the trend in the field of unregulated names being used in literature. It's difficult for scientists to review and compare data and communicate effectively because of this. Authors say that the SeqCode embraces findability, accessibility, interoperability, and reusability principles.
Chlamydia is an example. These organisms are unable to be named because they can't be grown, stored, or distributed as pure cultures.
It could be difficult for clinicians to remember the names of newly discovered STDs. There is a risk of those names being poorly cataloged, which could stifle tracking of disease outbreak and communication among scientists, doctors, and the public.
The controversy is over.
The move is not without controversy despite its intended goal of creating clarity and synergy.
The SeqCode follows a previous attempt by scientists to modify the ICNP to allow uncultivated prokaryotes to be named based on having a DNA sequence that would serve as the proof of their existence.
In 2020, a team led by Desert Research Institute biologist Alison Murray published a paper, also in Nature Microbiology, that was co-authored or endorsed by nearly 120 scientists representing 22 countries calling for action on the proposed modifications of the ICNP to accept DNA sequence as types. Modifications were turned down by the International Committee on Systematics of Prokaryotes.
It is clear that the global community of scientists is ready for a paradigm change in how we name prokaryotes. Modern genome technologies can resolve genomes of uncultivated organisms at the high degree of precision needed to ensure integrity. The way to communicate their existence and evolutionary history is to name them.
The "alternative route" which led to the creation of the SeqCode was the result of the 2020 setbacks.
Many people came to the table to share their perspectives and skills. The response to our workshops from scientists all over the world was incredible, and helped confirm why the time has come to change the name of prokaryotes.
Some scientists argue that more can be known about uncultivated prokaryotes than about those that can be grown in a lab. It is also true of studies of pure cultures that there are nuances in processing and interpreting the data.
According to the authors, the new system is designed to improve communication between the scientific community and the Microbial Sciences.
"We view this 'SeqCode v.1.0' as a necessary first step towards a unified system of nomenclature to communicate the full diversity of prokaryotes and we will cooperate with the community towards the realization of this vision," authors wrote.
A paper titled "SeqCode: a nomenclatural code for prokaryotes described from sequence data" was published in September.
More information: Brian P. Hedlund et al, SeqCode: a nomenclatural code for prokaryotes described from sequence data, Nature Microbiology (2022). DOI: 10.1038/s41564-022-01214-9The SeqCode can be found at www.seqco.de.
Journal information: Nature Microbiology